top of page
RESEARCH
GENOMIC ARCHITECTURE & REGULATORY DYNAMICS
We study genomic regulatory logic, chromatin dynamics and cellular information flow in both normal physiology and pathological perturbations, through the joint integration of data-sets from a variety of complementary biological layers generated by high-throughput technologies:
DATA SCIENCE & AI
-
Dynamics of Homeostatic Dysregulation Driving Disease Progression
-
Computational methods for identifying TF direct targets (by integrating TF binding with gene expression: ChIP-seq & RNA-seq or microarrays) and studying their influence on disease phenotypes
-
Ex: Our study of the TP53 target gene networks in apoptosis and senescence, identified by integrating TF binding and gene expression changes:
-
Methods for Identifying upstream regulators of gene signatures (RNA-seq, microarrays)
-
Early Carcinogenesis of Renal cancer using single-cell omics (scRNA-seq, ATAC-seq) - Collaboration with the Vanharanta lab (UH)
-
Enhancer re-wiring in renal and pancratic cancer metastasis
-
-
Cancer Translatomics
-
Understanding translational potential through Ribosome profiling (Ribo-seq, TIS-seq, iCLIP) - Collaboration with Narita (CRUK Cambridge Institute) and Hodson labs (Stem Cell Institute)
-
Proteomics
-
-
Regulatory Dynamics of the Non-Coding (epi)Genome in Disease
-
Understanding combinatorial histone and chromatin modifications (Histone ChIP-seq, ChIP-exo, Cut&Tag, Cut&Run)
-
DNA Methylation based cancer early detection using AI (methylation arrays)
-
Regulatory potential of Enhancer-Promoter-Insulator interactions, Enhancer rewiring, chromatin accessibility, nucleosome dynamics and TF footprinting (ChIP-seq & ATAC-seq, scATAC-seq) - Basu lab (ICL)
-
Enhancer Promoter Interaction detection using AI (Machine and Deep learning)
-
-
Chromatin Dynamics
-
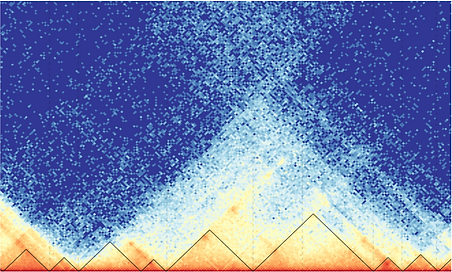
Transcriptional regulation via long-distance chromatin interactions and enhancer-promoter looping (Hi-C, micro-C, CHi-C and scHi-C)
-
Functional consequences of dynamic changes in chromatin 2D/3D structure: Hubs, Loops, TADs and A/B compartments (Hi-C, micro-C)
-
Functional genomics of DNA structural features such as iMotifs, G-quadruplexes, D-loops and R-loops
-
Effects of DNA-RNA hybrid structures such as R-loops through modulation of transcriptional regulation, and their contribution to genome instability and replicative stress (DRIP-seq)
-
- Lipidomics
- Integrating transcriptomics and lipidomics (LC-MS)
- NLRP3 Inflammasome (Collaborations with Anand Lab, ICL)
- Mitochondrial Lipids (Collaboration with Catala lab, ICL)

We utilize the latest advances in data science and artificial intelligence and apply these to biomedical research.

-
Artificial Intelligence methods for integrative 'Omics':
-
Machine Learning & Deep Learning for regulatory (epi)genomics
-
Piranha
-
-
Deep Learning for Biomedical Image analysis
-
linked Data, Knowledge Graphs and graph embedding based models for data integration, modelling and biomedical data mining
-
Senesceome-KG
-
-
ML and DL models for early detection and diagnosis of cancer
-
EMethylNET
-
MethylBoostER
-
-
-
Biomedical Natural Language Processing
-
NLP for extracting gene regulatory relationships from literature (PubMEd)
-
-
Biomedical Data Integration
-
Methods for statistical & computational biomedical data integration
-
SpiderSeqR
-
COBRA-tft
-
-
Deep Neural Graphs for regulatory epi-genomics and pathway bioinformatics
-
Web technologies & Integrative data resources
-
INTERFEROME: IFN regulated target gene resource
-
ReMOAT: Microarray re-annotation resource
-
ISGverse: Updated and curated Interferon Stimulated Gene Resource
-
InterferonScape: IFN target gene identification (STAT and IRF targets) and analysis of IFN networks in disease.
-
-
-
Information Visualisation for Biomedicine
-
Use of JavaScript, Processing, and Shiny to develop bespoke interactive data visualization
-
Biomedical Complex systems
We develop Systems Biology approaches to decipher the complexities of Immunity and Cancer.

-
Systems Immunology
-
Complex Systems analysis methods to understand immune and inflammatory responses in cancer and other diseases
-
Immune evasion mechanisms and modulation of adaptive immune responses
-
Pattern/Danger Recognition systems, Interferon and Interleukin pathways and networks
-
Metabolic remodelling, immuno-metabolic perturbations, secretomes, and micro-environment Interactions
-
-
Network Biology
-
Modelling cellular interactions and information flow
-
Reverse engineering gene regulatory networks
-
Building integrative and predictive regulatory models, knowledge graphs and network maps of disease processes - Senesceome-KG, InterferonScape
-
Developing resources for better pathway bioinformatics and linked data integration using graph-based strategies - Collaboration with Pietro Lio
-
-
Cancer Genomics & Computational Modelling
-
Data-driven quantitative (Mathematical/Statistical) modelling and inference
-
Impact of mutations (CNVs) and genetic aberrations (CNAs) on cancer pathways and networks (Whole Genome or Exome Sequencing)
-
Drug responses in cancer
-
bottom of page

